
pUNO1-SpikeV11
-
Cat.code:
p1-spike-v11
- Documents
Optimized SARS-CoV-2 Spike gene (Omicron – B.1.1.529/BA.1) for mammalian cell expression
pUNO1-SpikeV11 and pUNO1-SpikeV11-dfur plasmids have been specifically designed for the expression of the SARS-CoV-2 Spike (S) protein in mammalian cells with either a functional or inactivated furin (dfur) cleavage site. These plasmids encode the full-length Spike sequence from the Omicron variant (B.1.1.529/BA.1), and for optimal cellular expression, it is codon-optimized and the C‑terminal ER-retention signal has been removed [1, 2].
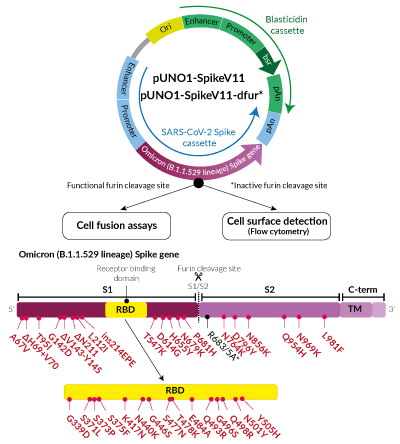
Omicron Variant (B.1.1.529/BA.1 lineage) Spike Expression vectors
for cell fusion and flow cytometry assays
Gene Description
These plasmids encode the Spike protein from the SARS-CoV-2 Omicron variant, first reported in South Africa in late November 2021. This variant is classified as a member of Clade 21K/B.1.1.529(BA.1) lineage (Nextstrain/Pango lineage classification). It is characterized by the presence of several mutations within the Spike coding region, of which, several are of concern [3].
- S1 domain: A67V, deletion (Δ)H69-V70, T95I, G142D, ΔV143-Y145, ΔN211, L212I, ins214EPE, T547K, D614G, H655Y, N679K, P681H
- RBD: G339D, S371L, S373P, S375F, K417N, N440K, G446S, S477N, T478K, E484A, Q493R, Q498R, N501Y, Y505H
- S2 domain: N764K, D796Y, N856K, Q954H, N969K, L981F
![]() Learn more about SARS-CoV-2 Variants
Learn more about SARS-CoV-2 Variants
The Spike protein contains a furin cleavage site that affects its cellular expression [4, in-house data]. Therefore, depending on your application InvivoGen offers:
- pUNO1-SpikeV11: with a functional furin cleavage site and recommended for Spike/ACE2 cell fusion assays
- pUNO1-SpikeV11-dfur: with an inactive furin (dfur) cleavage site for improved surface expression and detection (flow cytometry)
General Plasmid Description
These plasmids feature a potent mammalian expression cassette composed of the ubiquitous human EF1α-HTLV composite promoter and the SV40 polyadenylation (pAn) signal. The codon-optimized ORF includes the native SARS-CoV-2 Spike signal sequence. The plasmids are selectable with Blasticidin in both E. coli and mammalian cells (transient and stable transfection).
Applications
- Spike-mediated cell fusion assays with pUNO1-SpikeV11
- Cell surface detection by flow cytometry with pUNO1-SpikeV11-dfur
- Screening of SARS-CoV-2 inhibitors including small molecules, monoclonal antibodies, or convalescent plasma
References:
1. Johnson, M.C. et al. 2020. Optimized Pseudotyping Conditions for the SARS-COV-2 Spike Glycoprotein. J Virol 94.
2. Ou, X. et al. 2020. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat Commun 11, 1620.
3. https://www.who.int/en/activities/tracking-SARS-CoV-2-variants/
4. Coutard, B. et al. 2020. The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antiviral Res 176, 104742.
Specifications
pUNO1-SpikeV11
- Origin: Omicron variant (B.1.1.529/BA.1 lineage)
- Sequence Reference: EPI_ISL_6841980
- Codon Optimized
- ORF size: 3756 bp
- Sequencing primers:
- Forward HTLV 5’UTR: TGCTTGCTCAACTCTACGTC
- Reverse SV40 pAn: AACTTGTTTATTGCAGCTT - Quality Control:
- Plasmid construct is confirmed by restriction analysis and full‑length open reading frame (ORF) sequencing.
- After purification by ion-exchange chromatography, predominant supercoiled conformation is verified by electrophoresis.
pUNO1-SpikeV11-dfur
- Origin: Omicron variant (B.1.1.529/BA.1 lineage)
- Sequence Reference: EPI_ISL_6841980
- Codon Optimized and R683/5A mutations
- ORF size: 3756 bp
- Sequencing primers:
- Forward HTLV 5’UTR: TGCTTGCTCAACTCTACGTC
- Reverse SV40 pAn: AACTTGTTTATTGCAGCTT - Quality Control:
- Plasmid construct is confirmed by restriction analysis and full‑length open reading frame (ORF) sequencing.
- After purification by ion-exchange chromatography, predominant supercoiled conformation is verified by electrophoresis.
Contents
pUNO1-SpikeV11 and pUNO1-SpikeV11-dfur are provided as follows:
Please note: Each plasmid is sold separately.
- 20 μg of lyophilized DNA
- 2 x 1 ml Blasticidin at 10 mg/ml
![]() The product is shipped at room temperature.
The product is shipped at room temperature.
![]() Lyophilized DNA should be stored at -20 °C.
Lyophilized DNA should be stored at -20 °C.
![]() Resuspended DNA should be stored at -20 °C and is stable for up to 1 year.
Resuspended DNA should be stored at -20 °C and is stable for up to 1 year.
![]() Blasticidin is a harmful compound. Refer to the safety data sheet for handling instructions. Store Blasticidin at 4 °C or -20 °C for up to two years. The product is stable for 2 weeks at 37 °C.
Blasticidin is a harmful compound. Refer to the safety data sheet for handling instructions. Store Blasticidin at 4 °C or -20 °C for up to two years. The product is stable for 2 weeks at 37 °C.
![]() Avoid repeated freeze-thaw cycles.
Avoid repeated freeze-thaw cycles.
Details
Furin cleavage site in the SARS-CoV-2 Spike protein
A furin cleavage sequence (RRxR) is found within a polybasic cleavage site (681-PRRSR/SVA-688) at the boundary between the S1 and S2 domains (S1/S2) of the Spike protein [1]. Furin is enriched in the Golgi apparatus, where it functions to cleave proteins into their 'mature/active forms'. Specifically, it is suggested that cleavage at this site by furin pre-primes the SARS-CoV-2 S protein during its production. This allows further processing by cell surface host proteases (e.g. TMPRSS2) upon binding to ACE2, which ultimately facilitates viral-host membrane fusion [2,3].
► In a mammalian expression system (e.g. HEK293 cells), to maximize the surface expression of the S protein, the furin cleavage site in InvivoGen's pUNO1-SpikeV10-dfur has been inactivated (in-house data). The crucial recognition residues have been mutated (R683A and R685A) ensuring that the S protein is not cleaved by furin.
Furthermore, the S protein possesses cell-cell fusogenic activity and has been shown to trigger large syncytia formation (multi-nucleated cells). Notably, overexpression of an uncleavable S protein (mutated/inactivated furin cleavage site) has been shown to not induce cell-cell fusion, suggesting that cleavage at the multibasic site is a requirement for syncytia formation [3].
► To study cell-cell fusion by the SARS-CoV-2 spike, InvivoGen offers the pUNO1-SpikeV11 plasmid.
References:
1. Coutard, B. et al. 2020. The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antiviral Res 176, 104742.
2. Johnson, B.A. et al. 2021. Loss of furin cleavage site attenuates SARS-CoV-2 pathogenesis. Nature
3. Papa, G. et al. 2021. Furin cleavage of SARS-CoV-2 Spike promotes but is not essential for infection and cell-cell fusion. PLoS Pathog 17, e1009246.
DOCUMENTS
Documents
Plasmid Map and Sequence
Plasmid Sequence
Safety Data Sheet
Certificate of analysis
Need a CoA ?