pUNO1His-SARS2-S1
-
Cat.code:
p1his-cov2-s1
- Documents
SARS-CoV-2 Spike S1, codon-optimized & tagged in C-term
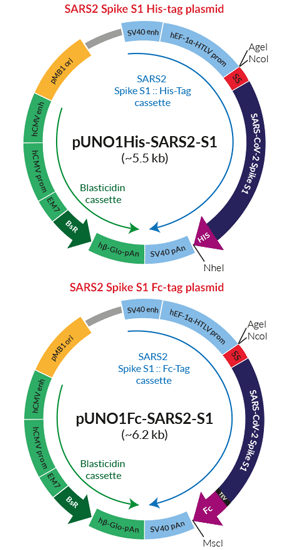
Schematic of tagged SARS2 Spike S1 domain production vectors
 InvivoGen also offers:
InvivoGen also offers:
• SARS-CoV-2 Cellular Receptor Genes
• SARS-CoV-2 Structural Genes
GENE DESCRIPTION
The SARS-CoV-2 (2019-nCoV) Spike S1 subunit plays a crucial role in the viral entry into the target cell. The S1 subunit features an N-term S1-NTD region and a C-term S1-CTD region. While S1-NTD is thought to mediate sugar-binding, the S1-CTD allows the virus to bind to ACE2 through the receptor-binding domain (RBD) [1-3]. In its resting conformation, S1 exerts a physical constraint on the Spike fusion subunit [3]. Research is ongoing to understand the exact mechanisms that drive conformation changes in S1 allowing subsequent membrane fusion events. S1 is a candidate for subunit vaccines against SARS-CoVs [4, 5].
pUNO1His-SARS2-S1 and pUNO1Fc-SARS2-S1 express a codon-optimized Spike S1 fused to a C-terminal His- or Fc-tag, respectively. The coding sequence is preceded by an exogenous signal sequence to ensure effective protein secretion. (See Details and Specifications for more information)
PLASMIDS DESCRIPTION
pUNO1His-SARS2-S1 and pUNO1Fc-SARS2-S1 are designed for the production in mammalian cells of the Spike S1 subunit. These vectors feature a potent mammalian expression cassette comprised of the strong SV40 enhancer, the ubiquitous human EF1α-HTLV composite promoter, and the SV40 polyadenylation (pAn) signal. The coding sequence is flanked by unique restriction sites, AgeI and NcoI at the 5' end, NheI at the 3' end of His-tag, and MscI at the 3' end of Fc-tag. Both plasmids are selectable with blasticidin in E. coli and mammalian cells.
QUALITY CONTROL
- Fully sequenced ORFs
- Predominant supercoiled conformation
![]() Learn more about SARS-CoV-2 infection cycle, immune responses, and potential therapeutics.
Learn more about SARS-CoV-2 infection cycle, immune responses, and potential therapeutics.
References
1. Li F., 2016. Structure, function, and evolution of coronavirus spike proteins. Annu. Rev. Virol. 3:237-261.
2. Li F. et al., 2005. Structure of SARS coronavirus spike receptor-binding domain complexed with receptor. Science. 309:1864-1868.
3. Walls A.C. et al., 2020. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell. 181(2):281-292.e6.
4. Wang N. et al., 2020. Subunit vaccines against emerging pathogenic human coronaviruses. Front. Microbiol. 11:298. DOI: 10.3389/fmicb.2020.00298.
5. Padron-Regalado E., 2020. Vaccines for SARS-CoV-2: Lessons from other coronavirus strains. Infect. Dis. Ther. DOI: 10.1007/s40121-020-00300-x.
Specifications
pUNO1His-SARS2-S1
- Strain: Wuhan-Hu-1 isolate
- ORF size (S1::His): 2097 bp
- Signal sequence: Exogenous (Lucia luciferase)
- Codon-optimized
- Tag: C-terminal 6xHistidine
-
Subcloning restriction sites: AgeI (in 5’) and NheI (in 3’)
- AgeI generates cohesive ends compatible with XmaI, BspEI, NgoMIV, and SgrAI restriction sites
- NheI generates cohesive ends compatible with AvrII, SpeI, and XbaI restriction sites -
Sequencing primers:
- Forward HTLV 5’UTR: TGCTTGCTCAACTCTACGTC
- Reverse SV40 pAn: AACTTGTTTATTGCAGCTT
pUNO1Fc-SARS2-S1
- Strain: Wuhan-Hu-1 isolate
- ORF size (S1::hIgG1 Fc): 2814 bp
- Signal sequence: Exogenous (Lucia luciferase)
- Codon-optimized
- Tag: C-terminal Human IgG1 Fc
-
Subcloning restriction sites: AgeI (in 5’) and MscI (in 3’)
- AgeI generates cohesive ends compatible with XmaI, BspEI, NgoMIV, and SgrAI restriction sites
- MscI generates blunt ends compatible with BaII, MIsI, MLuNI, Mox20I, and Msp20I restriction sites -
Sequencing primers:
- Forward HTLV 5’UTR: TGCTTGCTCAACTCTACGTC
- Reverse SV40 pAn: AACTTGTTTATTGCAGCTT
Contents
pUNO1His-SARS2-S1 and pUNO1Fc-SARS2-S1 contents:
- 20 μg of lyophilized DNA
- 2 x 1 ml Blasticidin at 10 mg/ml
![]() The product is shipped at room temperature.
The product is shipped at room temperature.
![]() Lyophilized DNA should be stored at -20 ̊C.
Lyophilized DNA should be stored at -20 ̊C.
![]() Resuspended DNA should be stored at -20 ̊C and is stable for up to 1 year.
Resuspended DNA should be stored at -20 ̊C and is stable for up to 1 year.
![]() Blasticidin is a harmful compound. Refer to the safety data sheet for handling instructions.
Blasticidin is a harmful compound. Refer to the safety data sheet for handling instructions.
Store blasticidin at 4°C or -20°C for up to two years. The product is stable for 2 weeks at 37°C.
Avoid repeated freeze-thaw cycles.
Details
Learn more about the SARS-CoV-2 Spike protein.
DOCUMENTS
Documents
Technical Data Sheet
Plasmid Map and Sequence
Safety Data Sheet
Plasmid Sequence
Certificate of analysis
Need a CoA ?


