pUNO1-SARS2-E
-
Cat.code:
puno1-cov2-e
- Documents
SARS-CoV-2 Envelope coding sequence
GENE DESCRIPTION
Envelope (E) is a small integral membrane protein, which together with Matrix/Membrane (M) and Spike(S) proteins, constitute coronaviruses’ interface to the external environment. The E protein is generally conserved across β-coronaviruses, and especially across SARS-CoVs. Although E is a minor viral component, it has demonstrated functions in virus assembly and release, and viral membrane curvature through its interaction with M [1].
pUNO1-SARS2-E contains the wild-type coding sequence of SARS-CoV-2 Envelope. (See Details and Specifications for more information).
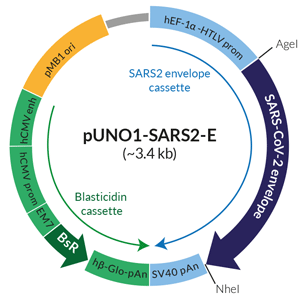
Schematic of SARS-CoV-2 envelope
expression vector
 InvivoGen also offers:
InvivoGen also offers:
• SARS-CoV-2-Cellular Receptor Genes
• SARS-CoV-2 S, S1, RBD, M, N Genes
PLASMID DESCRIPTION
pUNO1-SARS2-E features a potent mammalian expression cassette comprised of the ubiquitous human EF1α-HTLV composite promoter and the SV40 polyadenylation (pAn) signal. The ORF is flanked by unique restriction sites (AgeI and NheI) to facilitate its subcloning. The plasmid is selectable with blasticidin in both E. coli and mammalian cells. It can be used for transient or stable transfection. It contains no tag.
QUALITY CONTROL
- Fully sequenced ORF
- Predominant supercoiled conformation
![]() Learn more about SARS-CoV-2 infection cycle, immune responses, and potential therapeutics.
Learn more about SARS-CoV-2 infection cycle, immune responses, and potential therapeutics.
References
1. AJ Alsaadi E. & Jones I.M., 2019. Membrane binding proteins of coronaviruses. Future Virology. 14(4):275-286.
Specifications
- Strain: Wuhan-Hu-1 isolate
- Genbank: NC_045512.2
- ORF size from ATG to Stop codon: 228 bp
- Native (wild-type) sequence
-
Subcloning restriction sites in pUNO1: AgeI (in 5’) and NheI (in 3’)
- AgeI generates cohesive ends compatible with XmaI, BspEI, NgoMIV, and SgrAI restriction sites
- NheI generates cohesive ends compatible with AvrII, SpeI, and XbaI restriction sites -
Sequencing primers:
- Forward HTLV 5’UTR: TGCTTGCTCAACTCTACGTC
- Reverse SV40 pAn: AACTTGTTTATTGCAGCTT
Contents
- 20 μg of lyophilized DNA
- 2 x 1 ml blasticidin at 10 mg/ml
![]() The product is shipped at room temperature.
The product is shipped at room temperature.
![]() Lyophilized DNA should be stored at -20 ̊C.
Lyophilized DNA should be stored at -20 ̊C.
![]() Resuspended DNA should be stored at -20 ̊C and is stable up to 1 year.
Resuspended DNA should be stored at -20 ̊C and is stable up to 1 year.
![]() Blasticidin is a harmful compound. Refer to the safety data sheet for handling instructions.
Blasticidin is a harmful compound. Refer to the safety data sheet for handling instructions.
Store blasticidin at 4°C or -20°C for up to two years. The product is stable for 2 weeks at 37°C.
Avoid repeated freeze-thaw cycles.
Details
The current knowledge about the E protein comes from previous studies on β-coronaviruses other than SARS-CoV-2, including MERS-CoV and mouse hepatitis virus (MHV).
The CoV E protein is a short, integral membrane protein of 76-109 amino acids (a.a), and is comprised of three domains: a short hydrophilic N-terminal domain (7-12 a.a), a large hydrophobic transmembrane domain (25 a.a), and a long hydrophilic C-terminal domain [1]. The E protein interacts with multiple proteins form both the virus and the host.
In the viral particle, E interacts with the Matrix/Membrane (M) glycoprotein to induce the viral membrane curvature. The E protein also interacts with the Nucleocapsid (N) protein, independently of its interaction with M. The interaction of E with Spike (S) protein has not been extensively studied yet [1].
In the target cell, most of the E protein localizes to the ER and Golgi complex (ERGIC), where it participates in the assembly, budding and intracellular trafficking of the new virions. The fact that E is the least represented of the four structural proteins (M, S, N, and E), but is abundant in the ERGIC suggests that it exhibits other functions [1]. One of them is the formation of an ion-channel known as viroporin. Viroporin results from the CoV E protein oligomerization through homotypic interactions [1, 2]. Viroporin activity in mobilizing calcium ions and interacting with host tight-junction protein such as PALS1 (Proteins associated with Lin Seven 1) participates in the pathogenicity of some CoVs [1, 2]. Interestingly, the SARS-CoV E protein has been found to interact with four other host proteins: Bcl-xL, syntenin, sodium/phosphate (Na+/K+) ATPase alpha-1 subunit, and stomatin [1].
References
1. Schoeman D. & Fielding B.C., 2019. Coronavirus envelope protein: current knowledge. Virology Journal. 16:69.
2. AJ Alsaadi E. & Jones I.M., 2019. Membrane binding proteins of coronaviruses. Future Virology. 14(4):275-286.
DOCUMENTS
Documents
Technical Data Sheet
Plasmid Map and Sequence
Safety Data Sheet
Plasmid Sequence
Certificate of analysis
Need a CoA ?

