
pLV-SpikeV3
-
Cat.code:
plv-spike-v3
- Documents
For lentiviral particle pseudotyping with the SARS-CoV-2 Spike (B.1.351 - South African origin)
pLV-SpikeV3 has been specifically designed for the pseudotyping of lentiviral particles with the SARS-CoV-2 Spike (S) protein when co-transfected with plasmids encoding a reporter and accessory proteins (not provided). pLV-SpikeV3 encodes the full-length Spike sequence from the Beta variant (B.1.531), and for optimal expression, it is codon-optimized and the C‑terminal ER-retention signal has been removed [1, 2].
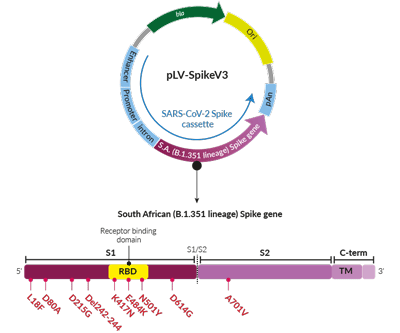
Beta Variant (B.1.351 lineage) Spike pseudotyping vector
Gene Description
This plasmid encodes the Spike protein from the SARS-CoV-2 Beta variant (B.1.351), first reported in South Africa in October 2020. This variant is classified as a member of Clade 20H/501Y.V2 and B.1.351 v2 lineage (Nextstrain/Pango lineage classification) [3]. It is characterized by the presence of a number of deletions and mutations within the Spike coding region, of which, several are of concern [3,4].
- S1 domain: L18F, D80A, D215G, deletion L242-L244, D614G
- RBD: K417N, E484K, N501Y
- S2 domain: A701V
![]() Learn more about SARS-CoV-2 variants
Learn more about SARS-CoV-2 variants
General Plasmid Description
This plasmid features a potent mammalian expression cassette composed of the ubiquitous human-(h)CMV composite promoter, a rabbit β-globin intron directly upstream of the spike gene, and a rabbit β-globin polyadenylation (pAn) signal. The spike coding sequence includes the SARS-CoV-2 signal sequence and the S1/S2 furin cleavage site [5]. These plasmids are selectable with ampicillin in E. coli.
Applications
- Generation of Spike pseudotyped lentiviral particles upon co-transfection with reporter and accessory protein plasmids (not provided).
- Screening of SARS-CoV-2 inhibitors including small molecules, monoclonal antibodies, or convalescent plasma.
References:
1. Johnson, M.C. et al. 2020. Optimized Pseudotyping Conditions for the SARS-COV-2 Spike Glycoprotein. J Virol 94.
2. Ou, X. et al. 2020. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat Commun 11, 1620.
3. Garcia-Beltran, W.F. et al. 2021. Multiple SARS-CoV-2 variants escape neutralization by vaccine-induced humoral immunity. Cell. doi:10.1016/j.cell.2021.03.013
4. Tegally, H. et al. 2020. Emergence and rapid spread of a new severe acute respiratory syndrome-related coronavirus 2 (SARS-CoV-2) lineage with multiple spike mutations in South Africa. Medrxiv. doi:10.1101/2020.12.21.20248640v1
5. Coutard, B. et al. 2020. The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antiviral Res 176, 104742.
Specifications
- Origin: Beta Variant (B.1.351 lineage) | South African origin
- Sequence Reference: GISAID EPI_ISL_745146
- Codon Optimized
- ORF size: 3756 bp
- Sequencing primers:
- Forward rbt β-globin intron: TGGTTACAATGATATACACTG
- Reverse rbt β-globin pAn: CTCAAGGGGCTTCATGATGTC - Quality Control:
- Plasmid construct is confirmed by restriction analysis and full‑length open reading frame (ORF) sequencing.
- After purification by ion-exchange chromatography, predominant supercoiled conformation is verified by electrophoresis.
Contents
pLV-SpikeV3 is provided as follows:
- 20 μg of lyophilized DNA
![]() The product is shipped at room temperature.
The product is shipped at room temperature.
![]() Lyophilized DNA should be stored at -20 °C.
Lyophilized DNA should be stored at -20 °C.
![]() Resuspended DNA should be stored at -20 °C and is stable up to 1 year.
Resuspended DNA should be stored at -20 °C and is stable up to 1 year.
Details
Spike-pseudotyping of lentiviral particles
InvivoGen's pLV-Spike plasmid collection has been designed for pseudotyping lentiviral particles with the SARS-CoV-2 Spike (S) protein. The S protein a structural glycoprotein expressed on the surface of SARS‑CoV-2. It mediates membrane fusion and viral entry into target cells upon binding to the host receptor ACE2 and its cleavage by cellular proteases such as TMPRSS2 [1]. Notably, the C‑terminal cytoplasmic tail of the S protein encodes a presumptive endoplasmic reticulum (ER)‑retention motif (KxHxx), which has previously been shown to enable the accumulation of SARS‑CoV S proteins at the ER‑Golgi intermediate compartment (ERGIC) and facilitate their incorporation into new virions [2]. The removal of this motif (d19) has been shown to increase the expression of the spike protein in pseudovirions [3,4].
Pseudotyped particle production involves the co-transfection of 293T cells with a reporter protein vector (e.g. GFP), one or several plasmids encoding the necessary lentiviral proteins, and the pseudotyping pLV-Spike plasmids. The transfected cells produce SARS‑CoV‑2 Spike (S)‑pseudotyped lentiviral particles, which can then be used to infect permissive cells, such as ACE2‑expressing HEK293-derived cells and ACE2‑TMPRSS2‑expressing A549-derived cells.
References
1. Hoffmann M. et al. 2020. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell. 181:1-16.
2. Ujike, M. et al. 2016. The contribution of the cytoplasmic retrieval signal of severe acute respiratory syndrome coronavirus to intracellular accumulation of S proteins and incorporation of S protein into virus-like particles. J Gen Virol 97, 1853-1864.
3. Johnson, M.C. et al. 2020. Optimized pseudotyping conditions for the SARS-COV2 Spike glycoprotein. J Virol. 94(21):e01062-20
4. Ou, X. et al. 2020. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat Commun 11, 1620.
DOCUMENTS
Documents
Plasmid Sequence
Safety Data Sheet
Certificate of analysis
Need a CoA ?