RBD-LuciaV4 (B.1.429)
-
Cat.code:
rbd-lucia-v4
- Documents
ABOUT
Recombinant RBD fusion protein (B.1.429 variant - Californian origin) for ELISA & LIPS
RBD-LuciaV4 (B.1.429) (~52 kDa) is a soluble fusion protein composed of the Spike Receptor Binding Domain (RBD) from the SARS-CoV-2 Epsilon variant (B.1.429) fused to a C‑terminal Lucia luciferase tag. RBD-LuciaV4 (B.1.429) has been specifically designed to assess the binding affinity of anti-Spike antibodies using either ELISA or LIPS (luciferase immunoprecipitation systems) assays [1-3].
SARS-CoV-2 Spike RBD
RBD-LuciaV4 (B.1.429) contains the Spike RBD domain, including the receptor-binding motif (RBM) from the SARS-CoV-2 Epsilon variant, first reported in the California USA in September 2020. [4]. This variant is classified as a member of Clade 21C / B.1.429 lineage (Nextstrain/Pango lineage classification). It is characterized by the presence of a key mutation within the Spike RBD coding region, which is of concern [4].
- L452R
![]() Learn more about the emerging SARS-CoV-2 variants around the world
Learn more about the emerging SARS-CoV-2 variants around the world
Applications
Luciferase-tagged RBD proteins are ideal for studying the binding of anti-spike monoclonal antibodies (mAbs) by solid-phase ELISA and/or solution‑phase LIPS assays, as well as anti‑spike polyclonal antibodies in the sera of recovered COVID‑19 patients and/or vaccinees by LIPS [1-3].
- ELISA: the Lucia luciferase tag provides a larger dynamic range than the commonly used HRP detection.
- LIPS: for the detection of antibodies, against both linear and conformational epitopes.
Importantly, using InvivoGen's expanding collection of Spike variant RBD-Lucia proteins, it can be seen that the SARS-CoV-2 variants display varying binding affinities to the different clinically relevant anti-Spike mAbs (see right). RBD‑LuciaV4 (B.1.429) has been generated by recombinant DNA technology, produced in CHO cells, and purified by IMAC (Immobilized Metal Affinity Chromatography) using a C‑terminal histidine tag. Protein size and purity (>90%) have been validated by SDS‑PAGE and the absence of endotoxin contamination has been confirmed using cellular assays.
References:
1. Burbelo, P.D. et al. 2010. Antibody-profiling technologies for studying humoral responses to infectious agents. Expert Rev Vaccines 9, 567-578.
2. Haljasmagi, L. et al. 2020. LIPS method for the detection of SARS-CoV-2 antibodies to spike and nucleocapsid proteins. Eur J Immunol 50, 1234-1236.
3. Liang, Y. et al. 2021. A luciferase immunosorbent assay for quantitative detection of IgG antibodies against SARS-CoV-2 nucleoprotein. J Virol Methods 292, 114141.
4. Deng, X. et al. 2021. Transmission, infectivity, and antibody neutralization of an emerging SARS-CoV-2 variant in California carrying a L452R spike protein mutation. medRxiv doi: 10.1101/2021.03.07.21252647
All InvivoGen products are for internal research use only, and not for human or veterinary use.
InvivoGen also offers:
SPECIFICATIONS
Specifications
GISAID EPI_ISL_753546
ELISA, LIPS
Each lot is functionally tested and validated.
CONTENTS
Contents
-
Product:RBD-LuciaV4 (B.1.429)
-
Cat code:rbd-lucia-v4
-
Quantity:50 µg
- 1.5 ml of endotoxin-free water,
- 1 tube of QUANTI-Luc™ 4 Reagent
Shipping & Storage
- Shipping method: Room temperature
- -20°C
- Avoid repeated freeze-thaw cycles
Storage:
Caution:
Details
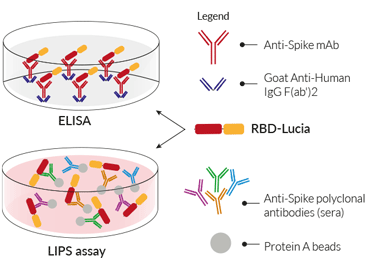
RBD-Lucia fusion protein for ELISA & LIPS
RBD-Lucia in ELISA
RBD-Lucia proteins can be used in a luciferase-based ELISA. Unlike a conventional ELISA, the plate is coated overnight with an Anti-human IgG F(ab')2 fragment. Upon addition of anti-spike monoclonal antibodies (mAb), they will bind to this 'capture' fragment through their Fc region, and RBD-Lucia will bind to the variable region. The luciferase activity is then used to assess the mAb binding affinity to the Spike RBD.
RBD-Lucia in LIPS
Currently, to perform a LIPS assay, soluble crude cell lysates or culture media of the luciferase tagged recombinant protein are extracted from transfected cells and directly used for the assay. InvivoGen's RBD-Lucia proteins streamline the protocol even further. Simply add the RBD-Lucia protein to either anti-spike mAbs or to anti‑spike polyclonal antibodies in the sera of recovered COVID‑19 patients and/or a vaccinee. Following this, antibody-protein complexes are purified using Protein A beads. Quantification of either binding affinity (mAb) and/or antibody levels (sera) is easily determined by assessing the Lucia luciferase activity.
DOCUMENTS
Documents
Technical Data Sheet
Validation Data Sheet
Safety Data Sheet
Certificate of analysis
Need a CoA ?







