Spotlight on COVID-19: Infection
Spotlight on COVID-19
• The infection cycle of SARS-CoV-2
• Treatment with repurposed drugs (updated)
COVID-19-RELATED PRODUCTS
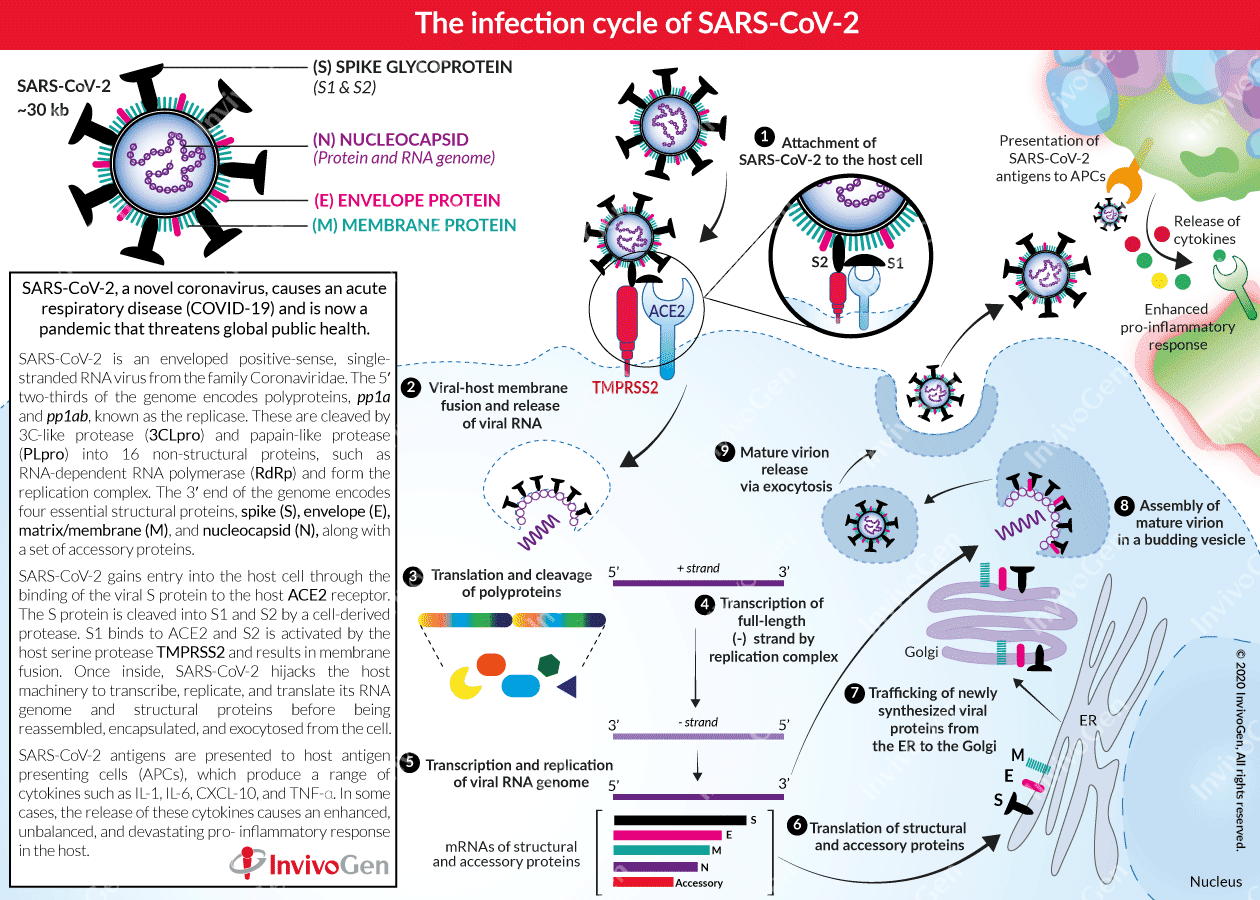
The infection cycle of SARS-CoV-2
Global Pandemic
Since December 2019, coronavirus disease 2019 (COVID-19) has spread rapidly around the world, causing a pandemic that threatens global public health. The causative agent of COVID-19 is Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2), a novel β-coronavirus. Globally, the number of positive COVID-19 cases grows exponentially every day and now the race is on to find both an effective treatment and to develop a vaccine for prevention measures in the future.
What is SARS-CoV-2?
Coronaviruses are relatively large enveloped, single-stranded, positive-sense RNA viruses (~30 kb). Their membrane has a crown-like appearance, due to its decoration with glycoprotein ‘spikes’ [1]. In particular, the β-coronavirus family includes Severe Acute Respiratory Syndrome (SARS) virus (SARS-CoV), Middle East respiratory syndrome (MERS) virus (MERS-CoV), and the COVID-19 causative agent SARS-CoV-2 [2, 3]. The 5′ two-thirds of the SARS-CoV-2 genome encodes two polyproteins, pp1a and pp1ab, collectively termed the replicase. These polyproteins are cleaved into 16 non-structural proteins including RNA-dependent RNA polymerase (RdRp) by two essential viral proteases, 3C-like protease (3CLpro) and papain-like protease (PLpro). In the 3′ one-third of the SARS-CoV-2 genome, like other β-coronavirus, encodes four essential structural proteins and a set of accessory proteins, which can all interfere with the host innate immune response [4, 5]. These structural proteins are [6]:
- Spike (S) glycoproteins – represent the largest structures of the virus and are essential for the entry into host cells.
- Small envelope (E) proteins - are only present in small quantities and most likely function as ion channels, not necessarily needed for viral replication but essential for pathogenesis.
- Membrane/Matrix (M) proteins – are the most abundant proteins in the virus structure and are responsible for viral membrane curvature and binding to the nucleocapsid.
- Nucleocapsid (N) proteins – bind to the viral RNA genome and ensure the maintenance of the RNA in a ‘beads-on-a-string’ conformation.
The SARS-CoV-2 infection cycle
Angiotensin-converting enzyme 2 (ACE2) is a cellular receptor expressed in the lungs, arteries, heart, kidneys, and the intestine. ACE2 binds to the viral (S) protein and constitutes the cellular entry receptor for SARS-CoV-2 into their human host [7]. More specifically, the S protein is cleaved into two subunits, S1 and S2, by an extracellular protease. While S1 binds to ACE2, S2 is further cleaved and activated by the host surface-associated transmembrane protease serine 2 (TMPRSS2) [8]. Together these actions result in host-viral membrane fusion and the release of the RNA genome into the host cell cytoplasm. Firstly, the host translational machinery is hijacked for the translation of the polyproteins and the essential viral proteases. The polyproteins (pp1a and pp1ab) are cleaved into 16 non-structural effector proteins by 3CLpro and PLpro allowing them to form the replication complex together with the RNA-dependent RNA polymerase, which synthesizes a full-length negative RNA strand template [1, 6]. This is used to replicate the complete RNA genome and generate the individual sub-genomic mRNA templates needed for the translation of the viral structural and accessory proteins. The newly synthesized structural and accessory proteins are then trafficked from the ER through the Golgi apparatus, after which new virions assemble in budding Golgi vesicles [6]. Finally, the mature SARS-CoV-2 virions are exocytosed and released from the host cell into the surrounding environment to repeat the infection cycle [2].
General host consequences
SARS-CoV-2 antigens are also presented to tissue residing antigen-presenting cells (APCs) such as macrophages, which in turn can produce a range of pro-inflammatory cytokines including IL‐1, IL‐4, IL‐6, IL‐8, MCSF, CXCL-10, and TNF-α [2, 9]. In some cases, these cytokines proliferate an enhanced, unbalanced, and devastating pro-inflammatory response in the host.
References
1. Liu, C. et al. 2020. Research and Development of Therapeutic Agents and Vaccines for COVID-19 and Related Human Coronavirus Diseases. ACS Central Science 6, 315-331.
2. Guo, Y.R. et al. 2020. The origin, transmission and clinical therapies on coronavirus disease 2019 (COVID-19) outbreak - an update on the status. Mil Med Res 7, 11.
3. Yin, Y. & Wunderink, R.G. 2018. MERS, SARS and other coronaviruses as causes of pneumonia. Respirology 23, 130-137.
4. Cui, J. et al. 2019. Origin and evolution of pathogenic coronaviruses. Nat Rev Microbiol 17, 181-192.
5. Zhu, N. et al. 2020. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N Engl J Med 382, 727-733.
6. Fehr, A.R. & Perlman, S. 2015. Coronaviruses: an overview of their replication and pathogenesis. Methods Mol Biol 1282, 1-23.
7. Zhou, P. et al. 2020. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270-273.
8. Hoffmann, M. et al. 2020. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell
9. Li, G. et al. 2020. Coronavirus infections and immune responses. J Med Virol 92, 424-432.



