The Online Tool to Choose and Design short hairpin RNAs
Based on the research from various laboratories including our own, InvivoGen has developed siRNA Wizard, a free software accessible online, that will help you find the best siRNA sequences on your target gene.
The siRNA Wizard tool will also design the pair of oligonucleotides needed to generate shRNAs using InvivoGen‘s psiRNA plasmids.
Select Candidate siRNA/shRNAs
The siRNA Wizard algorithm allows to select effective and specific siRNAs/shRNAs against your gene of interest based on thermodynamic and sequence-related criteria. Two search options are available:
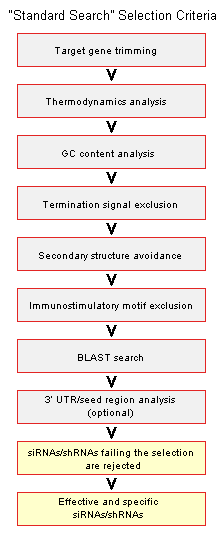
- Standard Search:
“Standard Search” uses default criteria described in the “siRNA/shRNA Design Guidelines” paragraph to select several candidate siRNAs/shRNAs against your gene of interest.
- Advanced Search:
“Advanced Search” lets you manually set the selection criteria.
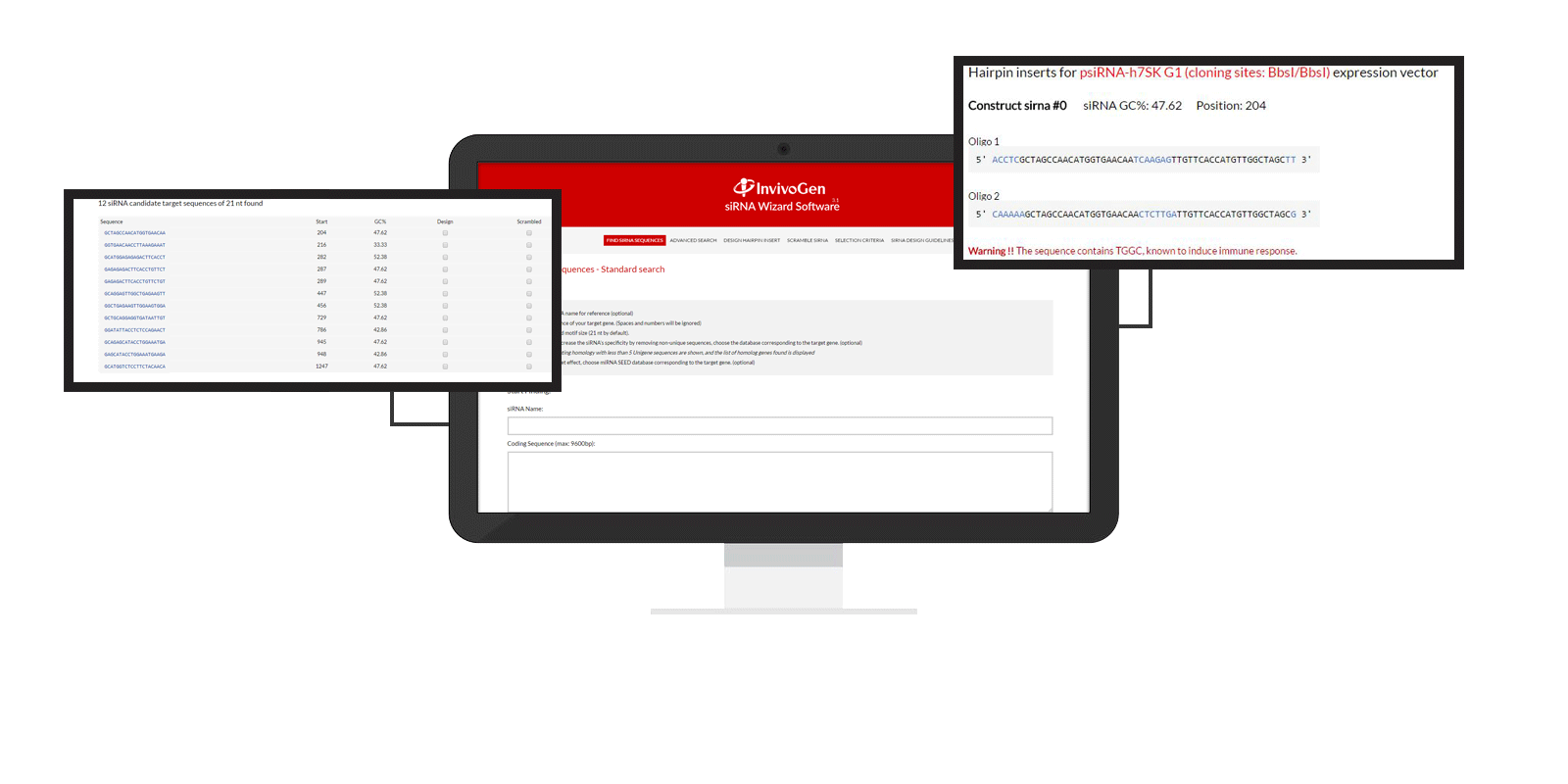
Design Hairpin Insert
Using your selected siRNA/shRNA sequence, this tool will design two complementary oligonucleotides necessary to create the hairpin insert for psiRNA cloning vectors and let you choose the sequence of the loop.
Generate siRNA/shRNA Scramble Sequence
This tool will return a scramble sequence with no match with any mRNA of the selected species database. This scramble sequence that serves as a negative control contains the same nucleotide composition as the selected siRNA/shRNA sequence.